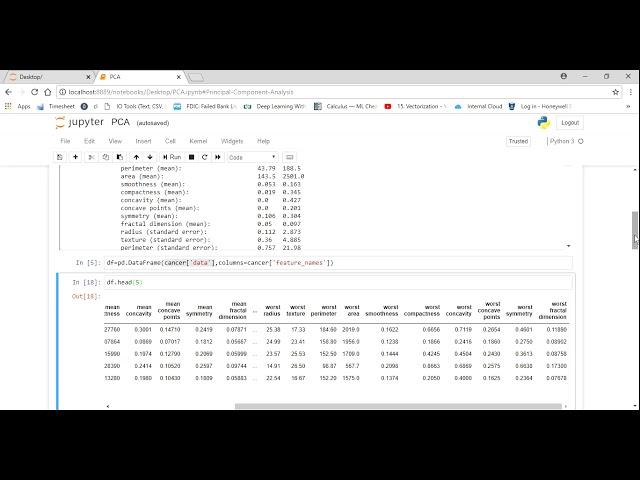
Principle Component Analysis (PCA) using sklearn and python
Комментарии:

Well explained it with an easy to understand example. Thanks
Ответить
I have my own data with some column of questionnaire, so what will be my column name there on the code
for instance, you put columns = cancer[feature_name], what I will put there on my own data?
all the column name one by one?
df-pd.DataFrame( cancer data' columns-cancer ['feature _nanes")

really sir, thanks for the knowledge. it helped me to solve your assignment in machine learning segment from PW skill..
Ответить
Actually !! I took a PCA on digit recognition data there I have took n_component value as 2 but in the visualization it coming multiple colors after executing. can anyone say what will be solution for that ?
Ответить
Nice Video
Ответить
so much insightful. The concept is well understood.
Ответить
amazing
simple and straightforward

So nice
Ответить
HELLO can you please make video on PCA use along with clustering and then explain the PCA values obtained in clusters
Ответить
can you please upload the data set
Ответить
Line 2 its a library that help you to import the dataset?
Ответить
thanks
Ответить
Hello, I have newly started working on a PCR project. I am stuck at a point and could really use some help...asap
Thanks a lot in advance.
I am working on python. So we have created PCA instance using PCA(0.85) and transformed the input data.
We have run a regression on principal components explaining 85 percent variance(Say N components). Now we have a regression equation in terms of N PCs. We have taken this equation and tried to express it in terms of original variables.
Now, In order to QC the coefficients in terms of original variables, we tried to take the N components(85% variance) and derived the new data back from this, and applied regression on this data hoping that this should give the same coefficients and intercept as in the above derived regression equation.
The issue here is that the coefficients are not matching when we take N components but when we take all the components the coefficients and intercept are matching exactly.
Also, R squared value and the predictions provided by these two equations are exactly same even if the coefficients are not matching
I am soo confused right now as to why this is happening. I might be missing out on the concept of PCA at some point. Any help is greatly appreciated.Thank you!

from sklearn.decompostition import PCA
pca=PCA()
pc=pca.fit_transform(df)
plt.figure()
plt.plot(np.cumsum(pca.explained_varience_ratio))
plt.xlabel('Column')
plt ylabel('EVR')
plt.show()

Excellent.. Appreciate it. .. liked your video
Ответить
Literally, I have searched and seen many videos , but this one has the best explanation
Ответить
Hi , in this scenerio we had 2 outputs, what happens when the number of outcome increases. For my case, I have 4 output
Ответить
Your video is very helpful. God bless you brother
Ответить
can we get to know what dimensions have been reduced and what 2 left there?
how we will infer from the graph after applying pca

how can we find the variance between the 2 components that the code is reduced to??
Ответить
i feel so good by seeing this ..thanks bro ...you help me out little bit ...make more videos on this type..
Ответить
A very insightful video
Ответить
great effort. thankyou!
Ответить
Thanks a lot Krish
Ответить
A lot of people in comment asked about intuition of PCA. So here it is
.We plot samples using the given features. for example imagine plotting different students (samples) on 3D graph (features English literature marks, Math marks and English Language Marks, x axis English literature marks ,y English Language and z axis Math ). Intuitively Someone who is good in English Literature must be good in English Language , so if I ask u to consider only two dimensions(features) for any classification model ,you will consider Maths and either of English, bcz we know by experience the variation in both English subjects would be less. Thus in PCA we actually project the samples (students in our example) in n numbers of PCA axis and choose the PCA which explains the maximum variation in data.
If we add variation of all PCAs it will be 1 or 100%.
Thus, instead of using all three subject marks I would rather use PC1 and PC2 as my features.
For PCA follow the steps
1. Once u have these 3d plot ready we calculate PC1 , which is a best fitting line that passes through the origin
2. Calculate Slope of PC1
3. calculate the eigen vector for the best fitting line
4.Find PC2 i.e. is a line perpendicular to PC1 and passes through the origin
5. Now rotate the graph such that PC1 is x axis and Pc2 is y axis , and project ur samples
Its kind tough to imagine ,Do read out more. Hope this helps

Great Video . At the end of the video u said lost of data ,I guess Its not 100 percent correct to phrase that its not "loss of data:, its actually the essence or rather the info of the data is not lost rather its converged into two dimensions
Ответить
How to know that how much data wo lost on decreasing dimensionality and how many components are best ??
Ответить
It is known that PCA causes loss of interpretability of the features. What is the alternative to PCA if we don't want to lose the interpretability? @Krish Naik. In case we have 40K features and we want to reduce dimension of the dataset without loosing the interpretability.
Ответить
how to choose the number of components in PCA?
Ответить
Hi krish , how Eigen values and Eigen vectors plays a role in capturing Principal components
Ответить
Bhai mere agar features independent honge to pca lagaye hi kyun.
Ответить
At time 10.28, when we do .. plt.scatter(x_pca[:,0] , .. shouldn't the second parameter here be target output column!! Why are we plotting it against x_pca[:,1] ?
Ответить
ValueError: Found array with 0 sample(s) (shape=(0, 372)) while a minimum of 1 is required by StandardScaler.
But there is no missing value

thank you for solving my question!
Ответить
I think small mistake on MinMaxscala here only used standard scaler
Ответить
Very clear and teach to the point. Thanks a lot.
Ответить
Great work... 4 thumbs for you. Greetings from a master student.
Ответить
How did you come to know that 'data' and 'feature_names' need to be considered for creating a dataframe from the file? Could you please explain
Ответить
Exactly half of the video was intro to data loading and explanation.... Where is the PCA???
Ответить
Very neat explanation.
Ответить
Insightful video.. Can we have a PCA vs LDA comparison video?
Much appreciated work!

Hi Naik, do we apply the pca only on training dataset, or the whole dataset(training+test)? some litterature advise to apply pca on training only, but in this case how to predict test set with the transformed data? waiting for your reply, thank you in advance
Ответить
Sir how to know what features(column names) are selected with pca ?
Ответить
Thanks
Ответить
Apply Basic PCA on the iris dataset.
• Describe the data set. Should the dataset been standardized?
• Describe the structure of correlations among variables.
• Compute a PCA with the maximum number of components
.• Compute the cumulative explained variance ratio. Determine the number of
componentskby your computed values.
• Print thekprincipal components directions and correlations of thekprincipal compo-
nents with the original variables. Interpret the contribution of the original variables into
the PC.
• Plot the samples projected into thekfirst PCs.
• Color samples by their species

sir how to figure out no. of prinicipal componenets to which we want to reduce the original dimension ??
Ответить
Firstly, Thanks for explaining PCA technique very clearly. Suppose, we do not know the features of a higher dimensional data. Is there any way to find the features and target within the data ? Is that possible by any chance. I am working with Hyperspectral raw data.
Ответить
Excellent work for begineers
Ответить
Shouldn't we validate how many PCA's are needed ?
Ответить



![ESO VSE - Sanity's Edge - Exarchanic Yaseyla HM first clear - safe strat - Fortüna [PS/EU] ESO VSE - Sanity's Edge - Exarchanic Yaseyla HM first clear - safe strat - Fortüna [PS/EU]](https://invideo.cc/img/upload/a2pRVFAyRkpFWnM.jpg)